Genetic testing companies and geneticists in general use centiMorgans (cM) to measure lengths of DNA that relatives share. You may have heard that DNA contains sequences of nucleotides—adenine, cytosine, guanine, and thymine, which are abbreviated as A, C, G, and T. One natural way to measure lengths of DNA is in terms of the number of nucleotides a segment of DNA contains. This is used in many contexts and is known as a sequence’s physical length. Physical lengths are measured in units of base pairs (bp) and give the number of nucleotides a sequence contains. So, for example, “GATTACA” is 7 bp.
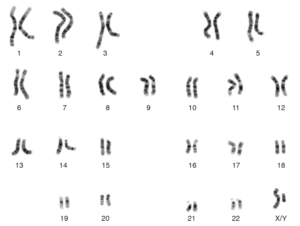
To understand centiMorgans, it’s useful to have a bit of background. We all have 23 pairs of chromosomes, having inherited one set of 23 from our father and another set of 23 from our mother. These chromosomes are physically small, with all 46 contained in our bodies’ cells, but they contain all of our DNA. The length of human chromosome 1 is roughly 249 million bp, whereas chromosome 22 is about 50.8 million bp.
When it comes to heredity, perhaps the most important cell types are the germ cells: sperm and eggs. While most human cells carry 23 pairs of chromosomes, germ cells contain only one copy of each chromosome. This is so that, once these cells fuse, the resulting fertilized egg will have 23 pairs of chromosomes.
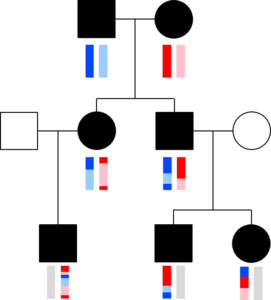 The chromosomes in germ cells are not simply an exact copy of one of the 23 chromosomes a person has, but are formed by recombination. A visualization helps capture this. The image with squares and circles shows how DNA from a couple might be transmitted to two children and three grandchildren. Here, circles represent females, squares represent males, and the vertical bars below these shapes give a colored representation of that person’s pair of chromosomes.1For simplicity, we will talk about recombination on only one chromosome. The same principles apply to all of them—chromosome 10, 2, etc.—except the X and Y chromosomes in fathers. At the top, the man has a dark and a light blue chromosome, and the woman has a red and a pink chromosome. Just below them are their two children, both of whom inherited one chromosome from each parent. Because of recombination, the children’s chromosomes are multi-colored, containing copies of DNA from the their mother’s two chromosomes and from their father’s chromosomes. In this case, both children received a copy of their dad’s dark blue chromosome at the top and both also received some amount of the light blue chromosome. Similarly, the mom transmitted a chromosome to each child containing some portions from her red chromosome and some from the pink chromosome. The bars get even more colorful in the next generation—for the shapes at the bottom—because these grandchildren inherited a chromosome that is recombined from their parents’ chromosomes. This means their chromosomes can contain pieces of all four of their grandparents’ chromosomes, and indeed, copies of DNA from all four chromosomes were transmitted to at least one grandchild.
The chromosomes in germ cells are not simply an exact copy of one of the 23 chromosomes a person has, but are formed by recombination. A visualization helps capture this. The image with squares and circles shows how DNA from a couple might be transmitted to two children and three grandchildren. Here, circles represent females, squares represent males, and the vertical bars below these shapes give a colored representation of that person’s pair of chromosomes.1For simplicity, we will talk about recombination on only one chromosome. The same principles apply to all of them—chromosome 10, 2, etc.—except the X and Y chromosomes in fathers. At the top, the man has a dark and a light blue chromosome, and the woman has a red and a pink chromosome. Just below them are their two children, both of whom inherited one chromosome from each parent. Because of recombination, the children’s chromosomes are multi-colored, containing copies of DNA from the their mother’s two chromosomes and from their father’s chromosomes. In this case, both children received a copy of their dad’s dark blue chromosome at the top and both also received some amount of the light blue chromosome. Similarly, the mom transmitted a chromosome to each child containing some portions from her red chromosome and some from the pink chromosome. The bars get even more colorful in the next generation—for the shapes at the bottom—because these grandchildren inherited a chromosome that is recombined from their parents’ chromosomes. This means their chromosomes can contain pieces of all four of their grandparents’ chromosomes, and indeed, copies of DNA from all four chromosomes were transmitted to at least one grandchild.
Considering all the chromosomes, a germ cell contains an average of 36.4 recombinations.2Technically, we should use the word crossover here. Strictly speaking, recombinations include both crossovers and another very small (10-100s of bp) form of recombination. We will follow this more typical use and say “recombination.” Said differently, there are an average of 36.4 recombinations per generation. In fact, this number is the Morgan length of all the chromosomes. That is, a Morgan is the average number of recombinations that occur in some piece of DNA in one generation. Of course, 36.4 Morgans is equal to 3640 cM: as its name implies, a centiMorgan is 1/100th of a Morgan.3Morgans are named for Thomas Hunt Morgan who led pioneering work in the study of recombination.
Researchers have analyzed DNA from many parents and children to measure how likely a region of DNA is to recombine in one generation. They have counted not just the average number of recombinations across the full genome—i.e., 3640 cM for all the chromosomes collectively—but in specific regions, like the average on chromosome 1, or some small section of chromosome 17. A 100 cM long section of DNA (which is the same as 1 Morgan long) will have, on average, 1 recombination per generation—so a parent will usually transmit one recombination in such a section. A piece of DNA with a length of 10 cM = 0.1 Morgans has a recombination in 1 out of 10 transmissions (10%). The parent-child DNA transmission data allow researchers to produce genetic maps that anyone can use to calculate the cM length of any physical span of DNA. Genetic testing companies use these maps to calculate the length of shared segments for relatives. Perhaps the most widely used genetic map measures chromosome 1 as 286 cM and chromosome 22 is 74.1 cM. It also shows the distance from chromosome 10 physical position 34,726,104 to 83,988,506 is 49.1 cM.
In an upcoming post, we’ll talk more about cM lengths of DNA and how recombination leads more distant relatives to share fewer segments that are also on average smaller than those that close relatives share.
DNA is a great way to solve family history questions.